Using Cutting-Edge Computation to Understand the Foundation's of Life's Mechanisms
Biological systems rely on the precise orchestration of molecular motions, interactions, and structures. In the Wereszczynski Lab, we leverage advanced computational approaches to unravel the physical principles underlying essential biological processes. By integrating molecular simulations, physical theory, and close collaboration with experimental teams, our research focuses on critical areas including gene regulation through chromatin dynamics, bacterial infection mechanisms, and methods for interpreting structural biology experiments. We aim to bridge physics and biology, translating atomic scale insights into a broader understanding of life itself.
Molecular Mechanisms of Epigenetic Regulation
The structure and motions of chromatin determine when and how genes are switched on or off. This control is fundamental to development, health, and disease. Our lab uses molecular simulations to understand how histone variants, chemical modifications, and nucleosome composition shape DNA packaging and gene accessibility. To get a deeper understanding of which physical mechanisms are truly essential, we also analyze simplified chromatin systems found in archaea and viruses. These comparisons let us pinpoint the rules that govern gene regulation across evolution and help address the central challenge of how cells control their genetic programs.
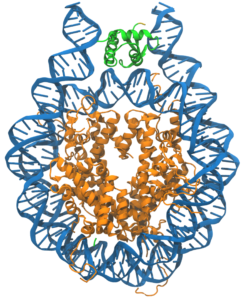
DNA (blue) is compacted by binding to eight core histones (orange), and further compacted through binding a linker histone (green) (adopted from Woods and Wereszczynski).
Molecular Underpinnings of Bacteria Virulence Pathway
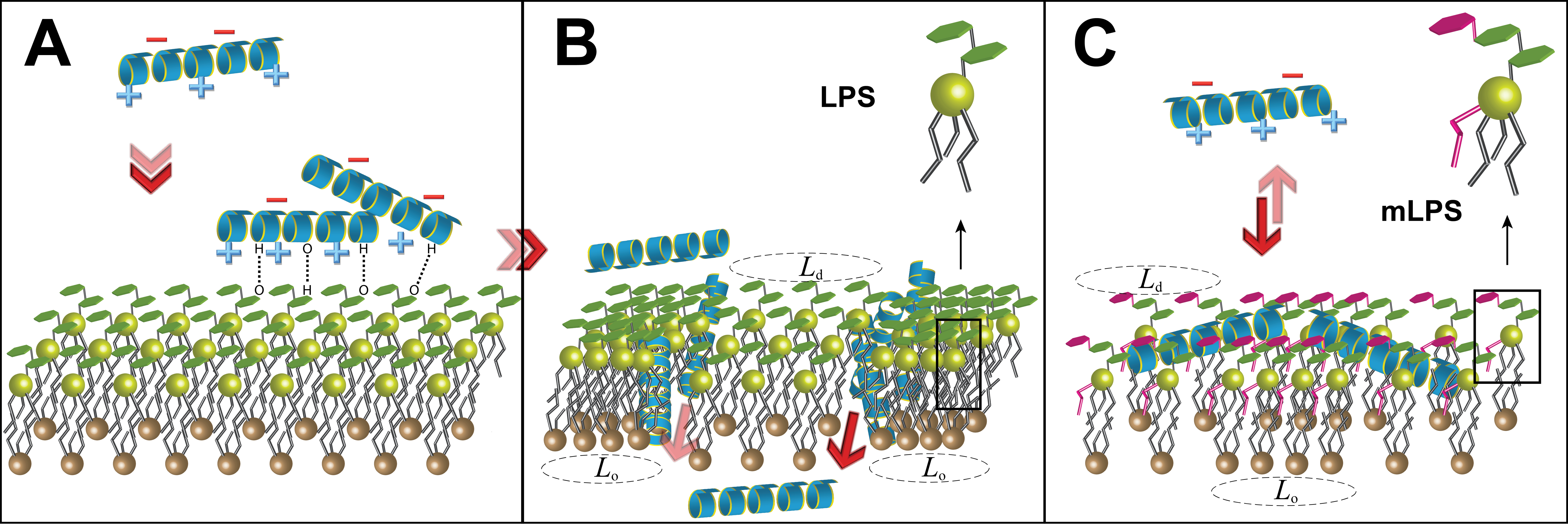
Mechanism of action of the antimicrobial peptide LL-37 against standard, LPS-containing, and modified LPS-containing bilayers (from Martynowycz et al.)
Antibiotic resistance is a major public health threat, and new approaches are urgently needed. Rather than focusing on killing bacteria, a promising strategy is to block virulence factors. These are molecular systems that allow bacteria to invade and harm their hosts. Disarming these pathways could slow the spread of resistance, since resistant strains must compete with bacteria that are still present but less harmful. The problem is that many virulence mechanisms remain poorly understood at the molecular level. Our lab uses molecular simulations and works closely with experimentalists to reveal how these systems function and how they can be disrupted by drugs or the immune system. By clarifying the molecular details of bacterial virulence, we help guide the development of new therapies that fight infection without driving resistance.
Robust Modelling of Small Angle Scattering Experiments
Small angle X-ray scattering (SAXS) is a powerful method for studying the structure and function of biomolecular complexes. SAXS stands out because it can analyze a wide range of targets, works under conditions close to those found in living cells, and captures information about flexible or disordered regions that are often missed by other techniques. However, the data produced by SAXS are low in resolution and provide only limited direct information about the detailed structures of proteins and complexes. Simulations can help bridge this gap by connecting scattering profiles to atomic-scale structures. In practice, this approach has been held back by limitations in physical modeling, the time and size of systems that can be simulated, and the complexity of the available software. As a result, a key challenge for the field is to develop methods that use SAXS data more effectively to interpret the structure, function, and dynamics of biomolecules.

Fitting of conventional and accelerated molecular dynamcis simulations results to experimental SAXS data (from Bowerman et al.).